 In other words, most of the court’s decisions are close to being in a two-dimensional subspace of all possible decisions. But Sirovich found that the third singular value is an order of magnitude smaller than the first one, so the matrix is well approximated by a matrix of rank 2. n Having developed this machinery, we complete our initial discussion of numerical linear algebra by deriving and making use of one final matrix factorization that exists for any matrix A 2 m n: the singular value decomposition (SVD). If the judges had made their decisions by flipping coins, this matrix would almost certainly have rank 9. In Chapter 5, we derived a number of algorithms for computing the eigenvalues and eigenvectors of matrices A 2 Rn. 
Since there are nine justices, each of whom takes a majority or minority position on each case, the data is a 468-by-9 matrix of +1s and -1s. His paper led to articles in the New York Times and the Washington Post because it provides a nonpolitical, phenomenological model of court decisions. Supreme Court" in the Proceedings of the US National Academy of Sciences. In 2003, Lawrence Sirovich of the Mount Sinai School of Medicine published "A pattern analysis of the second Rehnquist U.S.Tammy Kolda and Brett Bader, at Sandia National Labs in Livermore, ca, developed the Tensor Toolbox for MATLAB, which provides generalizations of PCA to multidimensional data sets.One example involves the analysis of the absorption spectrum of water samples from a lake to identify upstream sources of pollution. Rasmus Bro, a professor at the Royal Veterinary and Agricultural University in Denmark, and Barry Wise, head of Eigenvector Research in Wenatchee, Washington, both do chemometrics using SVD and PCA.The Wikipedia pages on SVD and PCA are quite good and contain a number of useful links, although not to each other.Professor SVD made all of this, and much more, possible. I came across some other interesting ones as I surfed around. Google finds over 3,000,000 Web pages that mention "singular value decomposition" and almost 200,000 pages that mention "SVD MATLAB." I knew about a few of these pages before I started to write this column. Eigenvalues are relevant when the matrix is regarded as a transformation from one space into itself-as, for example, in linear ordinary differential equations. Singular values are relevant when the matrix is regarded as a transformation from one space to a different space with possibly different dimensions. We can generate a 2-by-2 example by working backwards, computing a matrix from its SVD. By the time the first MATLAB appeared, around 1980, the SVD was one of its highlights. A variant of that algorithm, published by Gene Golub and Christian Reinsch in 1970 is still the one we use today. Kahan published the first effective algorithm in 1965. We did not yet have a practical way to actually compute it. A book that George Forsythe and I wrote in 1964 described the SVD as a nonconstructive way of characterizing the norm and condition number of a matrix. When I was a graduate student in the early 1960s, the SVD was still regarded as a fairly obscure theoretical concept. At the time, it had nothing to do with singular matrices. Picard used the adjective "singular" to mean something exceptional or out of the ordinary. Pete Stewart, author of the 1993 paper "On the Early History of the Singular Value Decomposition", tells me that the term valeurs singulières was first used by Emile Picard around 1910 in connection with integral equations. Let's validate the above with eigenfaces (i.e., the principal components / eigenvectors of the covariance matrix for such a face dataset) using the following face dataset: import numpy as npįrom sklearn.datasets import fetch_olivetti_faces The $j$-th principal component is given by $j$-th column of $\mathbf $ upto sign flip. Projections of the data on the principal axes are called principal components, also known as PC scores these can be seen as new, transformed, variables. The eigenvectors are called principal axes or principal directions of the data. 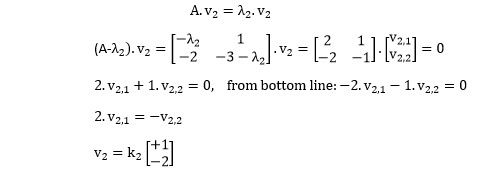
It is a symmetric matrix and so it can be diagonalized: $$\mathbf C = \mathbf V \mathbf L \mathbf V^\top,$$ where $\mathbf V$ is a matrix of eigenvectors (each column is an eigenvector) and $\mathbf L$ is a diagonal matrix with eigenvalues $\lambda_i$ in the decreasing order on the diagonal. 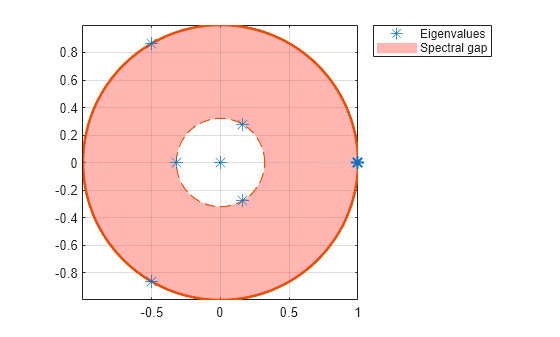
Then the $p \times p$ covariance matrix $\mathbf C$ is given by $\mathbf C = \mathbf X^\top \mathbf X/(n-1)$. column means have been subtracted and are now equal to zero. Let the real values data matrix $\mathbf X$ be of $n \times p$ size, where $n$ is the number of samples and $p$ is the number of variables. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
